Inspecting Demographics and Payment Patterns to Predict Credit Card Default Probability
Association:
iSchool, University of Toronto
Duration:
1.5 months
Data Science
Machine Learning
Python

For the final project in INF1340-Programming for Data Science, we worked on analyzing and predicting the default possibility of credit card owners.
Credit card debt and delinquency have always been one of the biggest pain points in the banking industry, implying socioeconomic and financial risks for both consumers and the servicing organizations. For consumers, delinquency can lead to a cycle of financial hardship, reduced access to credit, and long-term damage to creditworthiness. For financial institutions, high default rates increase operational risk, reduce profitability, and may necessitate stricter lending policies, which could limit access to credit for potential customers.
This study explores patterns and predictors of credit card default using a dataset that includes demographic, behavioral, and transactional variables. By employing descriptive and diagnostic analytics, such as pivot tables and statistical tests, and predictive modeling techniques, such as logistic regression and machine learning methods, the analysis identifies key factors contributing to default risk.
Diagnostic Analytics
After necessary data cleaning and descriptive analytics, we used diagnostic analytics to explore the potential relationships between variables, especially in relation to the target variable 'Default.'
# Visualize correlation with Heatmap def correlation_matrix(df): numerical_df = df.select_dtypes(include=np.number) plt.figure(figsize=(16, 14)) sns.heatmap(numerical_df.corr(), annot=True, cmap='coolwarm') plt.title('Correlation Heatmap') plt.show() return numerical_df.corr() correlation_matrix(credit_data)
# Visualize correlation with Heatmap def correlation_matrix(df): numerical_df = df.select_dtypes(include=np.number) plt.figure(figsize=(16, 14)) sns.heatmap(numerical_df.corr(), annot=True, cmap='coolwarm') plt.title('Correlation Heatmap') plt.show() return numerical_df.corr() correlation_matrix(credit_data)
# Visualize correlation with Heatmap def correlation_matrix(df): numerical_df = df.select_dtypes(include=np.number) plt.figure(figsize=(16, 14)) sns.heatmap(numerical_df.corr(), annot=True, cmap='coolwarm') plt.title('Correlation Heatmap') plt.show() return numerical_df.corr() correlation_matrix(credit_data)
# Visualize correlation with Heatmap def correlation_matrix(df): numerical_df = df.select_dtypes(include=np.number) plt.figure(figsize=(16, 14)) sns.heatmap(numerical_df.corr(), annot=True, cmap='coolwarm') plt.title('Correlation Heatmap') plt.show() return numerical_df.corr() correlation_matrix(credit_data)

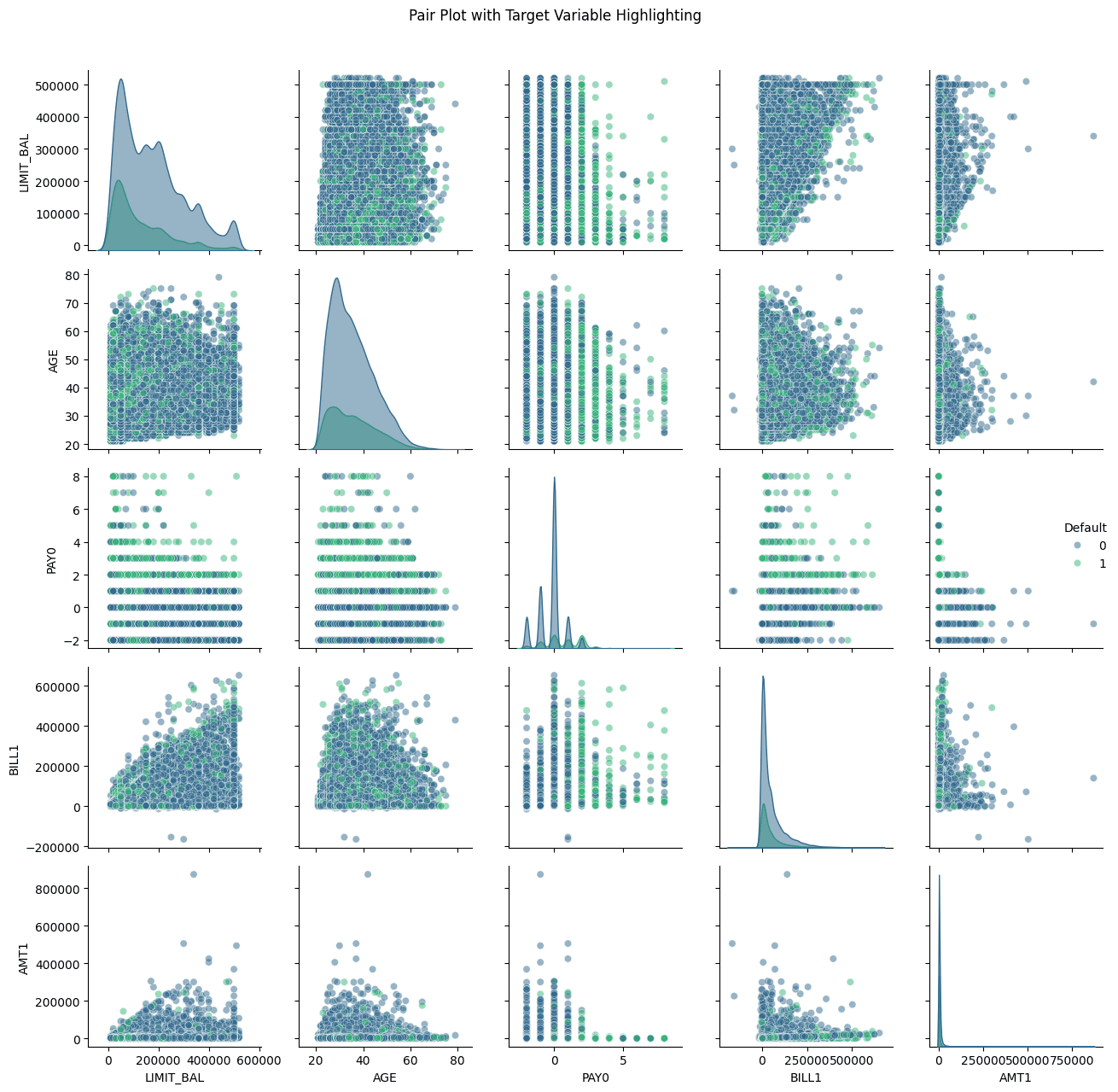
# Visualization for multivariate analysis def multivariate_analysis(df): # Pair Plot with Hue based on Target Variable plt.figure(figsize=(15,10)) subset_columns = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] g = sns.pairplot(df[subset_columns], hue='Default', plot_kws={'alpha': 0.5}, diag_kws={'alpha': 0.5}, palette='viridis') g.fig.suptitle('Pair Plot with Target Variable Highlighting', y=1.02) plt.tight_layout() plt.show() # Parallel Coordinates Plot using Plotly def normalize_column(series): return (series - series.min()) / (series.max() - series.min()) # Select columns for parallel coordinates parallel_cols = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] df_normalized = df[parallel_cols].copy() # Normalize numeric columns for col in parallel_cols[:-1]: # Exclude target variable df_normalized[col] = normalize_column(df_normalized[col]) # Create Parallel Coordinates Plot fig = px.parallel_coordinates( df_normalized, color='Default', title='Parallel Coordinates Plot', color_continuous_scale=px.colors.sequential.Viridis ) fig.show() # Box Plots for Categorical Variables categorical_columns = ['SEX', 'EDUCATION', 'MARRIAGE'] plt.figure(figsize=(15,5)) for i, var in enumerate(categorical_columns, 1): plt.subplot(1, 3, i) sns.boxplot(x=var, y='LIMIT_BAL', hue='Default', data=df, palette='Set2') plt.title(f'{var} vs Limit Balance by Default Status') plt.xticks(rotation=45) plt.tight_layout() plt.show() multivariate_analysis(credit_data)
# Visualization for multivariate analysis def multivariate_analysis(df): # Pair Plot with Hue based on Target Variable plt.figure(figsize=(15,10)) subset_columns = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] g = sns.pairplot(df[subset_columns], hue='Default', plot_kws={'alpha': 0.5}, diag_kws={'alpha': 0.5}, palette='viridis') g.fig.suptitle('Pair Plot with Target Variable Highlighting', y=1.02) plt.tight_layout() plt.show() # Parallel Coordinates Plot using Plotly def normalize_column(series): return (series - series.min()) / (series.max() - series.min()) # Select columns for parallel coordinates parallel_cols = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] df_normalized = df[parallel_cols].copy() # Normalize numeric columns for col in parallel_cols[:-1]: # Exclude target variable df_normalized[col] = normalize_column(df_normalized[col]) # Create Parallel Coordinates Plot fig = px.parallel_coordinates( df_normalized, color='Default', title='Parallel Coordinates Plot', color_continuous_scale=px.colors.sequential.Viridis ) fig.show() # Box Plots for Categorical Variables categorical_columns = ['SEX', 'EDUCATION', 'MARRIAGE'] plt.figure(figsize=(15,5)) for i, var in enumerate(categorical_columns, 1): plt.subplot(1, 3, i) sns.boxplot(x=var, y='LIMIT_BAL', hue='Default', data=df, palette='Set2') plt.title(f'{var} vs Limit Balance by Default Status') plt.xticks(rotation=45) plt.tight_layout() plt.show() multivariate_analysis(credit_data)
# Visualization for multivariate analysis def multivariate_analysis(df): # Pair Plot with Hue based on Target Variable plt.figure(figsize=(15,10)) subset_columns = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] g = sns.pairplot(df[subset_columns], hue='Default', plot_kws={'alpha': 0.5}, diag_kws={'alpha': 0.5}, palette='viridis') g.fig.suptitle('Pair Plot with Target Variable Highlighting', y=1.02) plt.tight_layout() plt.show() # Parallel Coordinates Plot using Plotly def normalize_column(series): return (series - series.min()) / (series.max() - series.min()) # Select columns for parallel coordinates parallel_cols = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] df_normalized = df[parallel_cols].copy() # Normalize numeric columns for col in parallel_cols[:-1]: # Exclude target variable df_normalized[col] = normalize_column(df_normalized[col]) # Create Parallel Coordinates Plot fig = px.parallel_coordinates( df_normalized, color='Default', title='Parallel Coordinates Plot', color_continuous_scale=px.colors.sequential.Viridis ) fig.show() # Box Plots for Categorical Variables categorical_columns = ['SEX', 'EDUCATION', 'MARRIAGE'] plt.figure(figsize=(15,5)) for i, var in enumerate(categorical_columns, 1): plt.subplot(1, 3, i) sns.boxplot(x=var, y='LIMIT_BAL', hue='Default', data=df, palette='Set2') plt.title(f'{var} vs Limit Balance by Default Status') plt.xticks(rotation=45) plt.tight_layout() plt.show() multivariate_analysis(credit_data)
# Visualization for multivariate analysis def multivariate_analysis(df): # Pair Plot with Hue based on Target Variable plt.figure(figsize=(15,10)) subset_columns = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] g = sns.pairplot(df[subset_columns], hue='Default', plot_kws={'alpha': 0.5}, diag_kws={'alpha': 0.5}, palette='viridis') g.fig.suptitle('Pair Plot with Target Variable Highlighting', y=1.02) plt.tight_layout() plt.show() # Parallel Coordinates Plot using Plotly def normalize_column(series): return (series - series.min()) / (series.max() - series.min()) # Select columns for parallel coordinates parallel_cols = ['LIMIT_BAL', 'AGE', 'PAY0', 'BILL1', 'AMT1', 'Default'] df_normalized = df[parallel_cols].copy() # Normalize numeric columns for col in parallel_cols[:-1]: # Exclude target variable df_normalized[col] = normalize_column(df_normalized[col]) # Create Parallel Coordinates Plot fig = px.parallel_coordinates( df_normalized, color='Default', title='Parallel Coordinates Plot', color_continuous_scale=px.colors.sequential.Viridis ) fig.show() # Box Plots for Categorical Variables categorical_columns = ['SEX', 'EDUCATION', 'MARRIAGE'] plt.figure(figsize=(15,5)) for i, var in enumerate(categorical_columns, 1): plt.subplot(1, 3, i) sns.boxplot(x=var, y='LIMIT_BAL', hue='Default', data=df, palette='Set2') plt.title(f'{var} vs Limit Balance by Default Status') plt.xticks(rotation=45) plt.tight_layout() plt.show() multivariate_analysis(credit_data)


# Logistic regression analysis def logistic_regression(df): # Select features and target variable X = X = df.drop(labels=['Default','AGE_GROUP'], axis=1) y = df['Default'] # Split data into training and test sets X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Logistic regression using statsmodels for detailed summary X_train_const = sm.add_constant(X_train) # Adding constant for intercept logit_model = sm.Logit(y_train, X_train_const) result = logit_model.fit() # Display the summary to interpret coefficients and p-values print(result.summary()) logistic_regression(credit_data) Logit Regression Results ============================================================================== Dep. Variable: Default No. Observations: 20858 Model: Logit Df Residuals: 20834 Method: MLE Df Model: 23 Date: Tue, 03 Dec 2024 Pseudo R-squ.: 0.1253 Time: 21:44:36 Log-Likelihood: -9613.9 converged: True LL-Null: -10991. Covariance Type: nonrobust LLR p-value: 0.000 ============================================================================== coef std err z P>|z| [0.025 0.975] ------------------------------------------------------------------------------ const -0.6448 0.144 -4.473 0.000 -0.927 -0.362 LIMIT_BAL -7.085e-07 1.93e-07 -3.663 0.000 -1.09e-06 -3.29e-07 SEX -0.1047 0.037 -2.830 0.005 -0.177 -0.032 EDUCATION -0.0685 0.026 -2.586 0.010 -0.120 -0.017 MARRIAGE -0.1851 0.038 -4.818 0.000 -0.260 -0.110 AGE 0.0054 0.002 2.524 0.012 0.001 0.010 PAY0 0.5908 0.021 27.797 0.000 0.549 0.632 PAY2 0.0832 0.024 3.427 0.001 0.036 0.131 PAY3 0.0560 0.027 2.056 0.040 0.003 0.109 PAY4 0.0511 0.030 1.696 0.090 -0.008 0.110 PAY5 0.0378 0.033 1.162 0.245 -0.026 0.102 PAY6 -0.0057 0.027 -0.213 0.832 -0.058 0.047 BILL1 -5.429e-06 1.39e-06 -3.918 0.000 -8.15e-06 -2.71e-06 BILL2 1.943e-06 1.85e-06 1.048 0.295 -1.69e-06 5.58e-06 BILL3 4.98e-07 1.68e-06 0.297 0.766 -2.79e-06 3.78e-06 BILL4 -1.174e-06 1.75e-06 -0.671 0.502 -4.6e-06 2.26e-06 BILL5 2.552e-06 1.97e-06 1.297 0.195 -1.31e-06 6.41e-06 BILL6 5.432e-07 1.47e-06 0.370 0.712 -2.34e-06 3.42e-06 AMT1 -1.216e-05 2.66e-06 -4.569 0.000 -1.74e-05 -6.95e-06 AMT2 -8.75e-06 2.46e-06 -3.552 0.000 -1.36e-05 -3.92e-06 AMT3 -5.074e-06 2.3e-06 -2.202 0.028 -9.59e-06 -5.59e-07 AMT4 -5.096e-06 2.19e-06 -2.327 0.020 -9.39e-06 -8.04e-07 AMT5 -3.515e-06 2.18e-06 -1.610 0.107 -7.79e-06 7.63e-07 AMT6 -1.779e-06 1.54e-06 -1.156 0.248 -4.8e-06 1.24e-06
# Logistic regression analysis def logistic_regression(df): # Select features and target variable X = X = df.drop(labels=['Default','AGE_GROUP'], axis=1) y = df['Default'] # Split data into training and test sets X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Logistic regression using statsmodels for detailed summary X_train_const = sm.add_constant(X_train) # Adding constant for intercept logit_model = sm.Logit(y_train, X_train_const) result = logit_model.fit() # Display the summary to interpret coefficients and p-values print(result.summary()) logistic_regression(credit_data) Logit Regression Results ============================================================================== Dep. Variable: Default No. Observations: 20858 Model: Logit Df Residuals: 20834 Method: MLE Df Model: 23 Date: Tue, 03 Dec 2024 Pseudo R-squ.: 0.1253 Time: 21:44:36 Log-Likelihood: -9613.9 converged: True LL-Null: -10991. Covariance Type: nonrobust LLR p-value: 0.000 ============================================================================== coef std err z P>|z| [0.025 0.975] ------------------------------------------------------------------------------ const -0.6448 0.144 -4.473 0.000 -0.927 -0.362 LIMIT_BAL -7.085e-07 1.93e-07 -3.663 0.000 -1.09e-06 -3.29e-07 SEX -0.1047 0.037 -2.830 0.005 -0.177 -0.032 EDUCATION -0.0685 0.026 -2.586 0.010 -0.120 -0.017 MARRIAGE -0.1851 0.038 -4.818 0.000 -0.260 -0.110 AGE 0.0054 0.002 2.524 0.012 0.001 0.010 PAY0 0.5908 0.021 27.797 0.000 0.549 0.632 PAY2 0.0832 0.024 3.427 0.001 0.036 0.131 PAY3 0.0560 0.027 2.056 0.040 0.003 0.109 PAY4 0.0511 0.030 1.696 0.090 -0.008 0.110 PAY5 0.0378 0.033 1.162 0.245 -0.026 0.102 PAY6 -0.0057 0.027 -0.213 0.832 -0.058 0.047 BILL1 -5.429e-06 1.39e-06 -3.918 0.000 -8.15e-06 -2.71e-06 BILL2 1.943e-06 1.85e-06 1.048 0.295 -1.69e-06 5.58e-06 BILL3 4.98e-07 1.68e-06 0.297 0.766 -2.79e-06 3.78e-06 BILL4 -1.174e-06 1.75e-06 -0.671 0.502 -4.6e-06 2.26e-06 BILL5 2.552e-06 1.97e-06 1.297 0.195 -1.31e-06 6.41e-06 BILL6 5.432e-07 1.47e-06 0.370 0.712 -2.34e-06 3.42e-06 AMT1 -1.216e-05 2.66e-06 -4.569 0.000 -1.74e-05 -6.95e-06 AMT2 -8.75e-06 2.46e-06 -3.552 0.000 -1.36e-05 -3.92e-06 AMT3 -5.074e-06 2.3e-06 -2.202 0.028 -9.59e-06 -5.59e-07 AMT4 -5.096e-06 2.19e-06 -2.327 0.020 -9.39e-06 -8.04e-07 AMT5 -3.515e-06 2.18e-06 -1.610 0.107 -7.79e-06 7.63e-07 AMT6 -1.779e-06 1.54e-06 -1.156 0.248 -4.8e-06 1.24e-06
# Logistic regression analysis def logistic_regression(df): # Select features and target variable X = X = df.drop(labels=['Default','AGE_GROUP'], axis=1) y = df['Default'] # Split data into training and test sets X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Logistic regression using statsmodels for detailed summary X_train_const = sm.add_constant(X_train) # Adding constant for intercept logit_model = sm.Logit(y_train, X_train_const) result = logit_model.fit() # Display the summary to interpret coefficients and p-values print(result.summary()) logistic_regression(credit_data) Logit Regression Results ============================================================================== Dep. Variable: Default No. Observations: 20858 Model: Logit Df Residuals: 20834 Method: MLE Df Model: 23 Date: Tue, 03 Dec 2024 Pseudo R-squ.: 0.1253 Time: 21:44:36 Log-Likelihood: -9613.9 converged: True LL-Null: -10991. Covariance Type: nonrobust LLR p-value: 0.000 ============================================================================== coef std err z P>|z| [0.025 0.975] ------------------------------------------------------------------------------ const -0.6448 0.144 -4.473 0.000 -0.927 -0.362 LIMIT_BAL -7.085e-07 1.93e-07 -3.663 0.000 -1.09e-06 -3.29e-07 SEX -0.1047 0.037 -2.830 0.005 -0.177 -0.032 EDUCATION -0.0685 0.026 -2.586 0.010 -0.120 -0.017 MARRIAGE -0.1851 0.038 -4.818 0.000 -0.260 -0.110 AGE 0.0054 0.002 2.524 0.012 0.001 0.010 PAY0 0.5908 0.021 27.797 0.000 0.549 0.632 PAY2 0.0832 0.024 3.427 0.001 0.036 0.131 PAY3 0.0560 0.027 2.056 0.040 0.003 0.109 PAY4 0.0511 0.030 1.696 0.090 -0.008 0.110 PAY5 0.0378 0.033 1.162 0.245 -0.026 0.102 PAY6 -0.0057 0.027 -0.213 0.832 -0.058 0.047 BILL1 -5.429e-06 1.39e-06 -3.918 0.000 -8.15e-06 -2.71e-06 BILL2 1.943e-06 1.85e-06 1.048 0.295 -1.69e-06 5.58e-06 BILL3 4.98e-07 1.68e-06 0.297 0.766 -2.79e-06 3.78e-06 BILL4 -1.174e-06 1.75e-06 -0.671 0.502 -4.6e-06 2.26e-06 BILL5 2.552e-06 1.97e-06 1.297 0.195 -1.31e-06 6.41e-06 BILL6 5.432e-07 1.47e-06 0.370 0.712 -2.34e-06 3.42e-06 AMT1 -1.216e-05 2.66e-06 -4.569 0.000 -1.74e-05 -6.95e-06 AMT2 -8.75e-06 2.46e-06 -3.552 0.000 -1.36e-05 -3.92e-06 AMT3 -5.074e-06 2.3e-06 -2.202 0.028 -9.59e-06 -5.59e-07 AMT4 -5.096e-06 2.19e-06 -2.327 0.020 -9.39e-06 -8.04e-07 AMT5 -3.515e-06 2.18e-06 -1.610 0.107 -7.79e-06 7.63e-07 AMT6 -1.779e-06 1.54e-06 -1.156 0.248 -4.8e-06 1.24e-06
# Logistic regression analysis def logistic_regression(df): # Select features and target variable X = X = df.drop(labels=['Default','AGE_GROUP'], axis=1) y = df['Default'] # Split data into training and test sets X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Logistic regression using statsmodels for detailed summary X_train_const = sm.add_constant(X_train) # Adding constant for intercept logit_model = sm.Logit(y_train, X_train_const) result = logit_model.fit() # Display the summary to interpret coefficients and p-values print(result.summary()) logistic_regression(credit_data) Logit Regression Results ============================================================================== Dep. Variable: Default No. Observations: 20858 Model: Logit Df Residuals: 20834 Method: MLE Df Model: 23 Date: Tue, 03 Dec 2024 Pseudo R-squ.: 0.1253 Time: 21:44:36 Log-Likelihood: -9613.9 converged: True LL-Null: -10991. Covariance Type: nonrobust LLR p-value: 0.000 ============================================================================== coef std err z P>|z| [0.025 0.975] ------------------------------------------------------------------------------ const -0.6448 0.144 -4.473 0.000 -0.927 -0.362 LIMIT_BAL -7.085e-07 1.93e-07 -3.663 0.000 -1.09e-06 -3.29e-07 SEX -0.1047 0.037 -2.830 0.005 -0.177 -0.032 EDUCATION -0.0685 0.026 -2.586 0.010 -0.120 -0.017 MARRIAGE -0.1851 0.038 -4.818 0.000 -0.260 -0.110 AGE 0.0054 0.002 2.524 0.012 0.001 0.010 PAY0 0.5908 0.021 27.797 0.000 0.549 0.632 PAY2 0.0832 0.024 3.427 0.001 0.036 0.131 PAY3 0.0560 0.027 2.056 0.040 0.003 0.109 PAY4 0.0511 0.030 1.696 0.090 -0.008 0.110 PAY5 0.0378 0.033 1.162 0.245 -0.026 0.102 PAY6 -0.0057 0.027 -0.213 0.832 -0.058 0.047 BILL1 -5.429e-06 1.39e-06 -3.918 0.000 -8.15e-06 -2.71e-06 BILL2 1.943e-06 1.85e-06 1.048 0.295 -1.69e-06 5.58e-06 BILL3 4.98e-07 1.68e-06 0.297 0.766 -2.79e-06 3.78e-06 BILL4 -1.174e-06 1.75e-06 -0.671 0.502 -4.6e-06 2.26e-06 BILL5 2.552e-06 1.97e-06 1.297 0.195 -1.31e-06 6.41e-06 BILL6 5.432e-07 1.47e-06 0.370 0.712 -2.34e-06 3.42e-06 AMT1 -1.216e-05 2.66e-06 -4.569 0.000 -1.74e-05 -6.95e-06 AMT2 -8.75e-06 2.46e-06 -3.552 0.000 -1.36e-05 -3.92e-06 AMT3 -5.074e-06 2.3e-06 -2.202 0.028 -9.59e-06 -5.59e-07 AMT4 -5.096e-06 2.19e-06 -2.327 0.020 -9.39e-06 -8.04e-07 AMT5 -3.515e-06 2.18e-06 -1.610 0.107 -7.79e-06 7.63e-07 AMT6 -1.779e-06 1.54e-06 -1.156 0.248 -4.8e-06 1.24e-06
Regarding the logistic regression, pseudo R-squared of 0.1253 indicates that about 12.53% of the variance in the dependent variable (Default) is explained by the model. The log-likelihood ratio (LLR) p-value is smaller than 0, which suggests the model is statistically significant overall. While pseudo R-squared values in logistic regression are typically lower than R-squared in linear regression, 12.53% suggests a moderate but not particularly strong fit.
Looking at the coefficients, p-values, and confidence intervals, the predictor’s impact on the likelihood of default is interpreted below:
LIMIT_BAL: The negative coefficient indicates that higher credit limits are associated with a lower likelihood of default. The coefficient is statistically significant (p < 0.05), but the effect size is small, suggesting that LIMIT_BAL alone may not be a strong predictor.
SEX, EDUCATION, MARRIAGE: These demographic variables are statistically significant, with p-values below 0.05. The signs of the coefficients suggest:
Male (coded as 1): Associated with a slightly lower likelihood of default.
Higher Education: Associated with a increased probability of default. Compared to 'Graduate School', having a 'High School' education is associated with a decrease in the log-odds of default by 0.1006.
Marital Status: The negative coefficient suggests that being married may reduce the likelihood of default.
AGE: Positively associated with default probability, meaning older age is associated with higher odds of default. Although significant, the small coefficient implies age alone has a limited effect on default likelihood.
Repayment History (PAY0 to PAY6): These are among the most influential predictors, as seen from large, positive coefficients for PAY0 and smaller yet significant coefficients for PAY2 and PAY3. The significance and positive signs indicate that higher values (i.e., delayed payments) increase the probability of default.
BILL1, BILL2, etc., and AMT1, AMT2, etc.: Many of these billing and payment amount features are statistically significant but with small coefficients, implying limited impact on default likelihood. Some values like BILL2 and AMT6 are not significant, meaning they don’t seem to add predictive value for default in this model.
Predictive Analytics
We used three different machine learning models for this dataset: Decision Tree, Logistic Regression, and Naive Bayes.
# Decision Tree def decision_tree(df): # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a basic Decision Tree clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Predict on test data and measure accuracy y_pred = clf.predict(X_test) accuracy = accuracy_score(y_test, y_pred) print(f'Basic Decision Tree Accuracy: {accuracy:.2f}') print(classification_report(y_test, y_pred)) # Experiment with different tree depths depths = [3, 5, 10, None] accuracies = [] for depth in depths: clf = DecisionTreeClassifier(criterion='entropy', max_depth=depth, random_state=42) clf.fit(X_train, y_train) y_pred = clf.predict(X_test) acc = accuracy_score(y_test, y_pred) accuracies.append(acc) print(f'Depth: {depth}, Accuracy: {acc:.2f}') # Plot accuracies vs. tree depths plt.figure(figsize=(10, 6)) plt.plot([str(d) for d in depths], accuracies, marker='o') plt.title('Decision Tree Accuracy vs. Depth') plt.xlabel('Tree Depth') plt.ylabel('Accuracy') plt.show() # Visualize the decision tree plt.figure(figsize=(20, 10)) tree.plot_tree(clf, feature_names=X.columns, class_names=['Yes', 'No'], filled=True) plt.title('Decision Tree Visualization') plt.show() decision_tree(credit_data) # Decision Tree Entropy def decision_tree_entropy(df): from sklearn.model_selection import train_test_split from sklearn.tree import DecisionTreeClassifier from sklearn.metrics import accuracy_score, classification_report from sklearn import tree # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a Decision Tree classifier using Information Gain (entropy) clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Extract feature importances (Information Gain) from the model information_gain = clf.feature_importances_ # Create a DataFrame to display feature names and their Information Gain feature_importance_df = pd.DataFrame({ 'Feature': X.columns, 'Information Gain': information_gain }).sort_values(by='Information Gain', ascending=False) # Display the DataFrame print(feature_importance_df) # Plot the Information Gain for each feature plt.figure(figsize=(12, 8)) plt.barh(feature_importance_df['Feature'], feature_importance_df['Information Gain'], color='skyblue') plt.xlabel('Information Gain') plt.ylabel('Feature') plt.title('Information Gain for Each Attribute') plt.gca().invert_yaxis() plt.show() decision_tree_entropy(credit_data) Basic Decision Tree Accuracy: 0.72 precision recall f1-score support 0 0.82 0.82 0.82 6918 1 0.38 0.39 0.39 2022 accuracy 0.72 8940 macro avg 0.60 0.60 0.60 8940 weighted avg 0.72 0.72 0.72 8940 Depth: 3, Accuracy: 0.82 Depth: 5, Accuracy: 0.81 Depth: 10, Accuracy: 0.80 Depth: None, Accuracy: 0.72
# Decision Tree def decision_tree(df): # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a basic Decision Tree clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Predict on test data and measure accuracy y_pred = clf.predict(X_test) accuracy = accuracy_score(y_test, y_pred) print(f'Basic Decision Tree Accuracy: {accuracy:.2f}') print(classification_report(y_test, y_pred)) # Experiment with different tree depths depths = [3, 5, 10, None] accuracies = [] for depth in depths: clf = DecisionTreeClassifier(criterion='entropy', max_depth=depth, random_state=42) clf.fit(X_train, y_train) y_pred = clf.predict(X_test) acc = accuracy_score(y_test, y_pred) accuracies.append(acc) print(f'Depth: {depth}, Accuracy: {acc:.2f}') # Plot accuracies vs. tree depths plt.figure(figsize=(10, 6)) plt.plot([str(d) for d in depths], accuracies, marker='o') plt.title('Decision Tree Accuracy vs. Depth') plt.xlabel('Tree Depth') plt.ylabel('Accuracy') plt.show() # Visualize the decision tree plt.figure(figsize=(20, 10)) tree.plot_tree(clf, feature_names=X.columns, class_names=['Yes', 'No'], filled=True) plt.title('Decision Tree Visualization') plt.show() decision_tree(credit_data) # Decision Tree Entropy def decision_tree_entropy(df): from sklearn.model_selection import train_test_split from sklearn.tree import DecisionTreeClassifier from sklearn.metrics import accuracy_score, classification_report from sklearn import tree # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a Decision Tree classifier using Information Gain (entropy) clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Extract feature importances (Information Gain) from the model information_gain = clf.feature_importances_ # Create a DataFrame to display feature names and their Information Gain feature_importance_df = pd.DataFrame({ 'Feature': X.columns, 'Information Gain': information_gain }).sort_values(by='Information Gain', ascending=False) # Display the DataFrame print(feature_importance_df) # Plot the Information Gain for each feature plt.figure(figsize=(12, 8)) plt.barh(feature_importance_df['Feature'], feature_importance_df['Information Gain'], color='skyblue') plt.xlabel('Information Gain') plt.ylabel('Feature') plt.title('Information Gain for Each Attribute') plt.gca().invert_yaxis() plt.show() decision_tree_entropy(credit_data) Basic Decision Tree Accuracy: 0.72 precision recall f1-score support 0 0.82 0.82 0.82 6918 1 0.38 0.39 0.39 2022 accuracy 0.72 8940 macro avg 0.60 0.60 0.60 8940 weighted avg 0.72 0.72 0.72 8940 Depth: 3, Accuracy: 0.82 Depth: 5, Accuracy: 0.81 Depth: 10, Accuracy: 0.80 Depth: None, Accuracy: 0.72
# Decision Tree def decision_tree(df): # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a basic Decision Tree clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Predict on test data and measure accuracy y_pred = clf.predict(X_test) accuracy = accuracy_score(y_test, y_pred) print(f'Basic Decision Tree Accuracy: {accuracy:.2f}') print(classification_report(y_test, y_pred)) # Experiment with different tree depths depths = [3, 5, 10, None] accuracies = [] for depth in depths: clf = DecisionTreeClassifier(criterion='entropy', max_depth=depth, random_state=42) clf.fit(X_train, y_train) y_pred = clf.predict(X_test) acc = accuracy_score(y_test, y_pred) accuracies.append(acc) print(f'Depth: {depth}, Accuracy: {acc:.2f}') # Plot accuracies vs. tree depths plt.figure(figsize=(10, 6)) plt.plot([str(d) for d in depths], accuracies, marker='o') plt.title('Decision Tree Accuracy vs. Depth') plt.xlabel('Tree Depth') plt.ylabel('Accuracy') plt.show() # Visualize the decision tree plt.figure(figsize=(20, 10)) tree.plot_tree(clf, feature_names=X.columns, class_names=['Yes', 'No'], filled=True) plt.title('Decision Tree Visualization') plt.show() decision_tree(credit_data) # Decision Tree Entropy def decision_tree_entropy(df): from sklearn.model_selection import train_test_split from sklearn.tree import DecisionTreeClassifier from sklearn.metrics import accuracy_score, classification_report from sklearn import tree # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a Decision Tree classifier using Information Gain (entropy) clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Extract feature importances (Information Gain) from the model information_gain = clf.feature_importances_ # Create a DataFrame to display feature names and their Information Gain feature_importance_df = pd.DataFrame({ 'Feature': X.columns, 'Information Gain': information_gain }).sort_values(by='Information Gain', ascending=False) # Display the DataFrame print(feature_importance_df) # Plot the Information Gain for each feature plt.figure(figsize=(12, 8)) plt.barh(feature_importance_df['Feature'], feature_importance_df['Information Gain'], color='skyblue') plt.xlabel('Information Gain') plt.ylabel('Feature') plt.title('Information Gain for Each Attribute') plt.gca().invert_yaxis() plt.show() decision_tree_entropy(credit_data) Basic Decision Tree Accuracy: 0.72 precision recall f1-score support 0 0.82 0.82 0.82 6918 1 0.38 0.39 0.39 2022 accuracy 0.72 8940 macro avg 0.60 0.60 0.60 8940 weighted avg 0.72 0.72 0.72 8940 Depth: 3, Accuracy: 0.82 Depth: 5, Accuracy: 0.81 Depth: 10, Accuracy: 0.80 Depth: None, Accuracy: 0.72
# Decision Tree def decision_tree(df): # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a basic Decision Tree clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Predict on test data and measure accuracy y_pred = clf.predict(X_test) accuracy = accuracy_score(y_test, y_pred) print(f'Basic Decision Tree Accuracy: {accuracy:.2f}') print(classification_report(y_test, y_pred)) # Experiment with different tree depths depths = [3, 5, 10, None] accuracies = [] for depth in depths: clf = DecisionTreeClassifier(criterion='entropy', max_depth=depth, random_state=42) clf.fit(X_train, y_train) y_pred = clf.predict(X_test) acc = accuracy_score(y_test, y_pred) accuracies.append(acc) print(f'Depth: {depth}, Accuracy: {acc:.2f}') # Plot accuracies vs. tree depths plt.figure(figsize=(10, 6)) plt.plot([str(d) for d in depths], accuracies, marker='o') plt.title('Decision Tree Accuracy vs. Depth') plt.xlabel('Tree Depth') plt.ylabel('Accuracy') plt.show() # Visualize the decision tree plt.figure(figsize=(20, 10)) tree.plot_tree(clf, feature_names=X.columns, class_names=['Yes', 'No'], filled=True) plt.title('Decision Tree Visualization') plt.show() decision_tree(credit_data) # Decision Tree Entropy def decision_tree_entropy(df): from sklearn.model_selection import train_test_split from sklearn.tree import DecisionTreeClassifier from sklearn.metrics import accuracy_score, classification_report from sklearn import tree # Select features and target variable X = df.drop(labels=['Default', 'AGE_GROUP'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Train a Decision Tree classifier using Information Gain (entropy) clf = DecisionTreeClassifier(criterion='entropy', random_state=42) clf.fit(X_train, y_train) # Extract feature importances (Information Gain) from the model information_gain = clf.feature_importances_ # Create a DataFrame to display feature names and their Information Gain feature_importance_df = pd.DataFrame({ 'Feature': X.columns, 'Information Gain': information_gain }).sort_values(by='Information Gain', ascending=False) # Display the DataFrame print(feature_importance_df) # Plot the Information Gain for each feature plt.figure(figsize=(12, 8)) plt.barh(feature_importance_df['Feature'], feature_importance_df['Information Gain'], color='skyblue') plt.xlabel('Information Gain') plt.ylabel('Feature') plt.title('Information Gain for Each Attribute') plt.gca().invert_yaxis() plt.show() decision_tree_entropy(credit_data) Basic Decision Tree Accuracy: 0.72 precision recall f1-score support 0 0.82 0.82 0.82 6918 1 0.38 0.39 0.39 2022 accuracy 0.72 8940 macro avg 0.60 0.60 0.60 8940 weighted avg 0.72 0.72 0.72 8940 Depth: 3, Accuracy: 0.82 Depth: 5, Accuracy: 0.81 Depth: 10, Accuracy: 0.80 Depth: None, Accuracy: 0.72


# Naıve Bayesian Classifiers def naive_bayes(df): # Select features and target variable X = df.drop(labels=['Default'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Scale the features scaler = StandardScaler() X_train_scaled = scaler.fit_transform(X_train) X_test_scaled = scaler.transform(X_test) # Model setup model = GaussianNB() model.fit(X_train_scaled, y_train) # Make predictions y_pred = model.predict(X_test_scaled) # Calculate metrics accuracy = accuracy_score(y_test, y_pred) report = classification_report(y_test, y_pred) conf_matrix = confusion_matrix(y_test, y_pred) # Print results print("\nNaive Bayesian Results:") print(f"Accuracy: {accuracy:.4f}") print("\nClassification Report:") print(report) # Plot confusion matrix plt.figure(figsize=(8, 6)) sns.heatmap(conf_matrix, annot=True, fmt='d', cmap='Blues') plt.title('Confusion Matrix - Naıve Bayesian') plt.ylabel('True Label') plt.xlabel('Predicted Label') plt.show() return { "model": model, "accuracy": accuracy, "classification_report": report, "confusion_matrix": conf_matrix, } results = naive_bayes(credit_data) Naive Bayesian Results: Accuracy: 0.7506 Classification Report: precision recall f1-score support 0 0.87 0.80 0.83 6918 1 0.46 0.58 0.51 2022 accuracy 0.75 8940 macro avg 0.66 0.69 0.67 8940 weighted avg 0.77 0.75
# Naıve Bayesian Classifiers def naive_bayes(df): # Select features and target variable X = df.drop(labels=['Default'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Scale the features scaler = StandardScaler() X_train_scaled = scaler.fit_transform(X_train) X_test_scaled = scaler.transform(X_test) # Model setup model = GaussianNB() model.fit(X_train_scaled, y_train) # Make predictions y_pred = model.predict(X_test_scaled) # Calculate metrics accuracy = accuracy_score(y_test, y_pred) report = classification_report(y_test, y_pred) conf_matrix = confusion_matrix(y_test, y_pred) # Print results print("\nNaive Bayesian Results:") print(f"Accuracy: {accuracy:.4f}") print("\nClassification Report:") print(report) # Plot confusion matrix plt.figure(figsize=(8, 6)) sns.heatmap(conf_matrix, annot=True, fmt='d', cmap='Blues') plt.title('Confusion Matrix - Naıve Bayesian') plt.ylabel('True Label') plt.xlabel('Predicted Label') plt.show() return { "model": model, "accuracy": accuracy, "classification_report": report, "confusion_matrix": conf_matrix, } results = naive_bayes(credit_data) Naive Bayesian Results: Accuracy: 0.7506 Classification Report: precision recall f1-score support 0 0.87 0.80 0.83 6918 1 0.46 0.58 0.51 2022 accuracy 0.75 8940 macro avg 0.66 0.69 0.67 8940 weighted avg 0.77 0.75
# Naıve Bayesian Classifiers def naive_bayes(df): # Select features and target variable X = df.drop(labels=['Default'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Scale the features scaler = StandardScaler() X_train_scaled = scaler.fit_transform(X_train) X_test_scaled = scaler.transform(X_test) # Model setup model = GaussianNB() model.fit(X_train_scaled, y_train) # Make predictions y_pred = model.predict(X_test_scaled) # Calculate metrics accuracy = accuracy_score(y_test, y_pred) report = classification_report(y_test, y_pred) conf_matrix = confusion_matrix(y_test, y_pred) # Print results print("\nNaive Bayesian Results:") print(f"Accuracy: {accuracy:.4f}") print("\nClassification Report:") print(report) # Plot confusion matrix plt.figure(figsize=(8, 6)) sns.heatmap(conf_matrix, annot=True, fmt='d', cmap='Blues') plt.title('Confusion Matrix - Naıve Bayesian') plt.ylabel('True Label') plt.xlabel('Predicted Label') plt.show() return { "model": model, "accuracy": accuracy, "classification_report": report, "confusion_matrix": conf_matrix, } results = naive_bayes(credit_data) Naive Bayesian Results: Accuracy: 0.7506 Classification Report: precision recall f1-score support 0 0.87 0.80 0.83 6918 1 0.46 0.58 0.51 2022 accuracy 0.75 8940 macro avg 0.66 0.69 0.67 8940 weighted avg 0.77 0.75
# Naıve Bayesian Classifiers def naive_bayes(df): # Select features and target variable X = df.drop(labels=['Default'], axis=1) y = df['Default'] # Train-test split X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42) # Scale the features scaler = StandardScaler() X_train_scaled = scaler.fit_transform(X_train) X_test_scaled = scaler.transform(X_test) # Model setup model = GaussianNB() model.fit(X_train_scaled, y_train) # Make predictions y_pred = model.predict(X_test_scaled) # Calculate metrics accuracy = accuracy_score(y_test, y_pred) report = classification_report(y_test, y_pred) conf_matrix = confusion_matrix(y_test, y_pred) # Print results print("\nNaive Bayesian Results:") print(f"Accuracy: {accuracy:.4f}") print("\nClassification Report:") print(report) # Plot confusion matrix plt.figure(figsize=(8, 6)) sns.heatmap(conf_matrix, annot=True, fmt='d', cmap='Blues') plt.title('Confusion Matrix - Naıve Bayesian') plt.ylabel('True Label') plt.xlabel('Predicted Label') plt.show() return { "model": model, "accuracy": accuracy, "classification_report": report, "confusion_matrix": conf_matrix, } results = naive_bayes(credit_data) Naive Bayesian Results: Accuracy: 0.7506 Classification Report: precision recall f1-score support 0 0.87 0.80 0.83 6918 1 0.46 0.58 0.51 2022 accuracy 0.75 8940 macro avg 0.66 0.69 0.67 8940 weighted avg 0.77 0.75

Overall, Naive Bayes (0.75) performs worse than both the linear regression (0.80) and decision tree models (up to 0.82). This lower accuracy compared to previous models suggests that Naive Bayes may not be the best choice for this dataset, especially given the complexity of relationships in financial data. Naive Bayes assumes feature independence, which might not hold for this dataset (e.g., financial metrics like billing amounts and payments are likely correlated). Naive Bayes achieves the highest recall for class 1 (0.58), indicating it is better at identifying defaults compared to the decision tree (0.39) and linear regression (0.24), but its precision for class 1 (0.46) is only slightly better than the decision tree (0.38) and worse than linear regression (0.70).
Conclusion
Key findings include:
Model-wise, the most important feature in predicting default status is PAY0–the most recent credit card payment status.
Younger age groups show higher default rates, likely reflecting financial inexperience or limited income.
Higher education levels are strongly associated with higher credit limits and might reduce the likelihood of default but their influence on default prediction is limited.
All models faced challenges due to the class imbalance (non-defaults significantly outnumber defaults).
This project demonstrates the trade-offs between different models in handling imbalanced datasets for credit default prediction. While logistic regression and decision trees excel in overall accuracy, Naive Bayes offers superior recall for defaults, which may align better with business goals focused on identifying risky customers. Further optimization through feature engineering, resampling, and advanced modeling techniques can enhance performance and address class imbalance effectively.
Other Projects
© 2026. All rights Reserved.
© 2026. All rights Reserved.

